 The clubber pipeline
The clubber pipeline
Microbial functional diversification is driven by environmental factors, i.e. microorganisms inhabiting the same environmental niche tend to be more functionally similar than those from different environments. In some cases, even closely phylogenetically related microbes differ more across environments than across taxa. While microbial similarities are often reported in terms of taxonomic relationships, no existing databases directly links microbial functions to the environment.
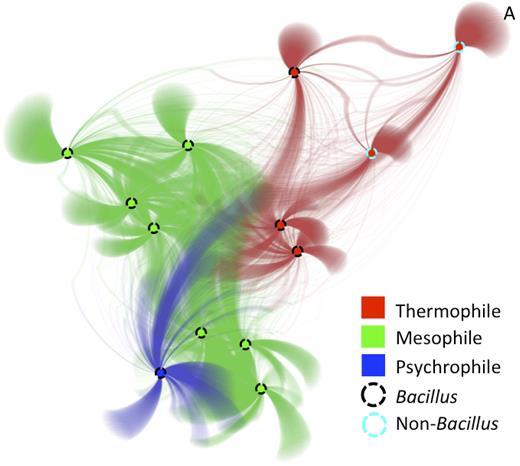
fusionDB is a database using functional data to represent 1,374 taxonomically distinct bacteria annotated with available metadata: habitat/niche, preferred temperature, and oxygen use. Each microbe is encoded as a set of functions represented by its proteome and individual microbes are connected via common functions. Users can search fusionDB via combinations of organism names and metadata (Explore). Moreover, the web interface allows mapping new microbial genomes to the functional spectrum of reference bacteria (Map Organism), rendering interactive similarity networks that highlight shared functionality.
fusionDB is a service that is provided free for academic use. If you use the service please cite:
- fusionDB: fusionDB: assessing microbial diversity and environmental preferences via functional similarity networks. Nucleic Acids Research
- Fusion: Functional Basis of Microorganism Classification. PLOS Computational Biology